Post-processing¶
Overview¶
pyRSKtools is RBR’s open source Python toolbox for reading, post-processing, visualizing, and exporting RBR logger data. Users may plot data as a time series or as depth profiles using tailored plotting utilities. Time-depth heat maps can be plotted easily to visualize transects or moored profiler data. A full suite of data post-processing functionality, such as methods to match sensor time constants and bin average, are available to enhance data quality.
You may review the pyRSKtools getting started guide for an introduction on how to connect to an RSK file via the RSK class.
pyRSKtools includes a series of methods to post-process RBR logger data. Below we show how to implement some common processing steps to obtain the highest quality data possible. All post-processing methods are customizable via input arguments. Documentation for each method can be accessed using the Python commands doc and help.
Basic Usage¶
To instantiate and open an RSK file, there are two approaches (see pyRSKtools getting started guide for more information). The context manager approach is shown below:
from pyrsktools import RSK
with RSK("/path/to/data.rsk") as rsk:
# Print a list of all the channels in the RSK file
rsk.printchannels()
# Read data
rsk.readdata()
# Derive sea pressure from total pressure
rsk.deriveseapressure()
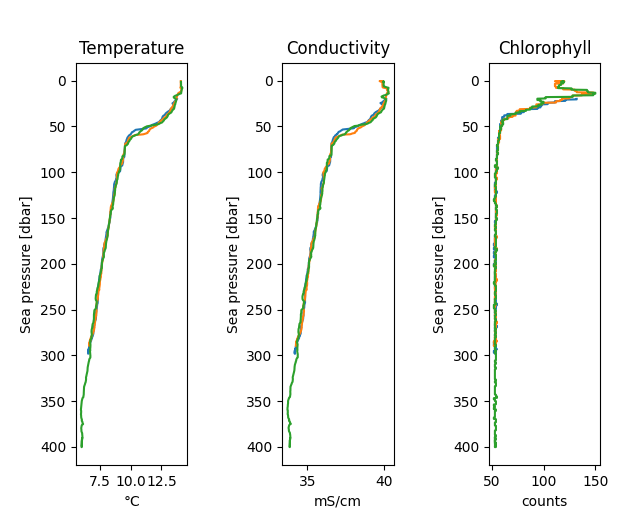
# Plot a few profiles of temperature, conductivity, and chlorophyll
fig, axes = rsk.plotprofiles(
channels=["conductivity", "temperature", "chlorophyll"],
profiles=range(0, 3),
direction="down",
)
plt.show()
Model: RBRconcerto³
Serial ID: 66098
Sampling period: 0.125 second
Channels: index name unit
_____ ____________________________ ________
0 conductivity mS/cm
1 temperature °C
2 pressure dbar
3 temperature1 °C
4 dissolved_o2_saturation %
5 backscatter counts
6 backscatter1 counts
7 chlorophyll counts
Derive sea pressure from total pressure¶
We suggest deriving sea pressure first, especially when an atmospheric pressure other than the nominal
value of 10.1325 dbar is desired. The patm argument is the atmospheric pressure used to calculate the sea pressure.
A custom value can be used; otherwise, the default is to retrieve the value stored in the parameters field
or to assume it is 10.1325 dbar if the parameter’s field is unavailable.
RSK.deriveseapressure() also supports a variable patm input as a list, when that happens,
the input list should have the same length as RSK.data. In this example, we’ll take atmospheric pressure to be 10 dbar.
rsk.deriveseapressure(patm = 10)
Data post-processing¶
What follows is a generic recipe for post-processing RBR CTD data. It is a guideline - RBR CTDs are used in many environments, with many sensor packages, and are profiled from a variety of vessels (or no vessel at all). While the basic approach here is relevant for many cases, the processing parameters may not apply widely.
First, keep a copy of the raw data to compare with the processed data.
raw = rsk.copy()
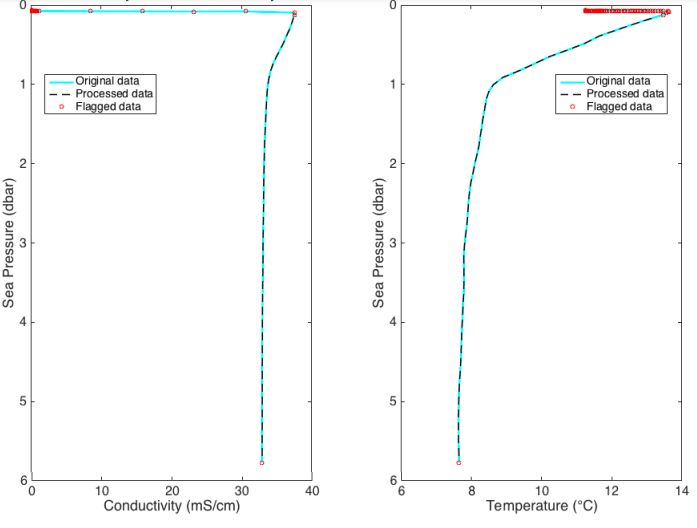
Correct for A2D zero-order hold¶
The analogue-to-digital (A2D) converter on RBR instruments must recalibrate periodically. In the time it
takes for the calibration to finish, one or more samples are missed. The instrument firmware fills the missed
sample with the same data measured during the previous sample, a technique called a zero-order hold.
RSK.correcthold() identifies zero-hold points by finding where consecutive differences of each channel
are equal to zero and then replaces these samples with a NaN or an interpolated value. See the RSK.correcthold() for further information.
rsk.correcthold(action = "interp")
Low-pass filtering¶
Low-pass filtering is commonly used to reduce noise and to match sensor time constants, typically for temperature and conductivity. Users may also wish to filter other channels to simply reduce noise (e.g., optical channels such as chlorophyll-a or turbidity).
Most RBR instruments designed for profiling are equipped with thermistors that have a time constant of 100 ms,
which is “slower” than the conductivity cell. When the time constants are different, salinity will contain spikes
at strong gradients. One solution is to “slow down” the conductivity sensor to match the thermistor. In this example dataset,
the logger sampled at 6 Hz (found in the ScheduleInfo class using `rsk.scheduleInfo.samplingperiod()),
so a 5 sample running average provides more than sufficient smoothing to match the time response of the conductivity sensor to the thermistor.
rsk.smooth(channels = ["salinity"], windowLength = 5)
Alignment of conductivity and temperature¶
Conductivity and temperature often need to be aligned in time to account for the fact these sensors are not always co-located on the logger. The implication is that, under dynamic conditions (e.g., profiling), the sensors are measuring a slightly different parcel of water at any instant.
Furthermore, sensors with long time constants introduce a time lag to the data. For example, dissolved oxygen sensors often have a long time constant, and this delays the measurement relative to the true value. This can be fixed to some degree by advancing the sensor data in time.
When temperature and conductivity are misaligned, salinity will contain spikes at sharp interfaces and a bias in continuously stratified environments. Properly aligning the sensors, together with matching the time response, will minimize salinity spiking and bias.
A common approach to determine the optimal lag is to compute and plot salinity for a range of lags and
choose the lag (often by eye) with the smallest salinity spikes at sharp temperature interfaces.
As an alternative approach, pyRSKtools includes a method called RSK.calculateCTlag() that estimates
the optimal lag between conductivity and temperature by minimizing salinity spiking. We currently suggest
using both approaches to check for consistency. See the API reference for RSK.calculateCTlag() for more
information.
As a rough guide, temperature from a CTD equipped with the red combined CT cell and a fast thermistor typically requires only a very small-time advance (perhaps tens of milliseconds). Temperature from a CTD equipped with a cylindrical black conductivity cell (with the thermistor on the sensor endcap) typically requires a temperature lag correction of about 0.1 s to 0.3 s (1 or 2 samples at 6 Hz).
import numpy as np
# Required shift of C relative to T for each profile
lag = rsk.calculateCTlag(seapressureRange = (1,50), direction = "down")
# Advance temperature
lag = -np.array(lag)
# Select best lag for consistency among profiles
lag = np.median(lag)
rsk.alignchannel(channel = "temperature", lag = lag, direction = "down")
Users wishing to learn more about dynamic sensor corrections and RBR CTDs are encouraged to watch a special RBR webinar on dynamic errors from May 2020 (Youtube and PDF).
Remove loops¶
Working in rough seas can cause the CTD profiling rate to vary, and even change signs (i.e., the CTD
momentarily changes direction). When this happens, the CTD effectively samples its own wake,
degrading the quality of the profile in regions of strong gradients. The measurements taken when the instrument
is profiling too slowly or during a pressure reversal should not be used for further analysis.
We recommend using RSK.removeloops() to flag and treat the data when the instrument (1) falls below a threshold speed and
(2) when the pressure reverses (the CTD “loops”). Before using RSK.removeloops(), use
RSK.deriveseapressure() to calculate sea pressure from total pressure, RSK.derivedepth() to calculate depth
from sea pressure, and then use RSK.derivevelocity() to calculate profiling rate.
In the example dataset, good data is collected on the upcast. RSK.removeloops(), when applied to this
data, removes data when the instrument is profiled slowly near the surface.
rsk.deriveseapressure()
rsk.derivedepth()
rsk.derivevelocity()
# Apply the algorithm
rsk = RSKremoveloops(threshold = 0.3)
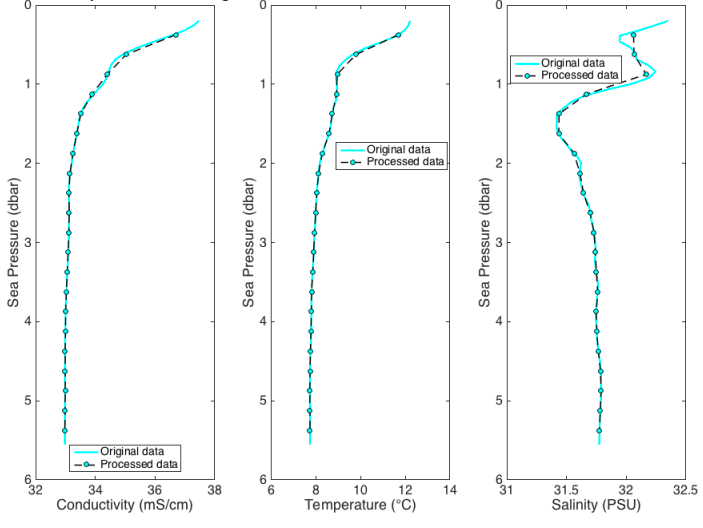
Derived variables¶
pyRSKtools includes a few convenience methods to derive common oceanographic variables. For
example, RSK.derivesalinity() computes Practical Salinity using the TEOS-10 GSW function
gsw_SP_from_C. RSK.derivesalinity() is a wrapper for the
TEOS-10 GSW function gsw_SP_from_C, and adds a new channel called "salinity" as a column
in RSK.data. The official Python implementation of the TEOS-10 GSW toolbox is freely available
and can be found here.
rsk.deriveseapressure()
rsk.derivedepth()
rsk.derivevelocity()
rsk.derivesalinity()
rsk.derivesigma()
# Print a list of channels in the rsk file
rsk.printchannels()
Model: RBRconcerto³
Serial ID: 66098
Sampling period: 0.125 second
Channels: index name unit
_____ ____________________________ ________
0 conductivity mS/cm
1 temperature °C
2 pressure dbar
3 temperature1 °C
4 dissolved_o2_saturation %
5 backscatter counts
6 backscatter1 counts
7 chlorophyll counts
8 sea_pressure dbar
9 depth m
10 velocity m/s
11 salinity PSU
12 density_anomaly kg/m³
Bin average all channels by sea pressure¶
Bin averaging reduces sensor noise and ensures that each profile is referenced to a common grid. The latter
is often an advantage for plotting data as “heatmaps.” RSK.binaverage() allows users to bin channels
according to any reference, but the most common choices are time, depth, and sea pressure. It also can
handle grids with a variable bin size. In the following example, the data are averaged into 0.25 dbar bins.
rsk.binaverage(
binBy = "sea_pressure",
binSize = 0.25,
boundary = [0.5, 5.5],
direction = "up"
)
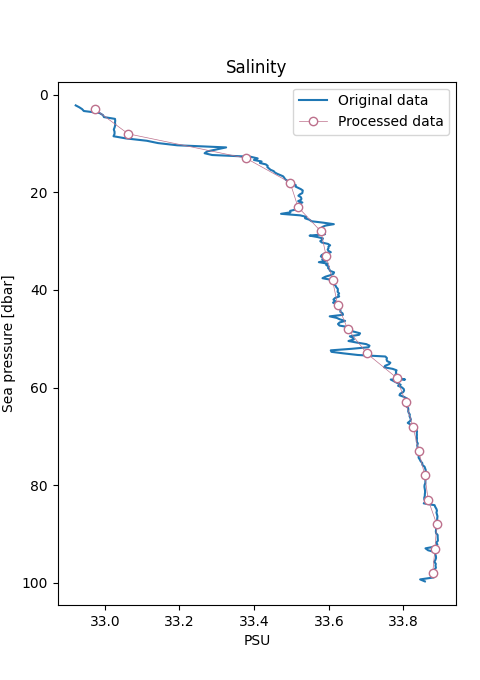
Compare the raw and processed data¶
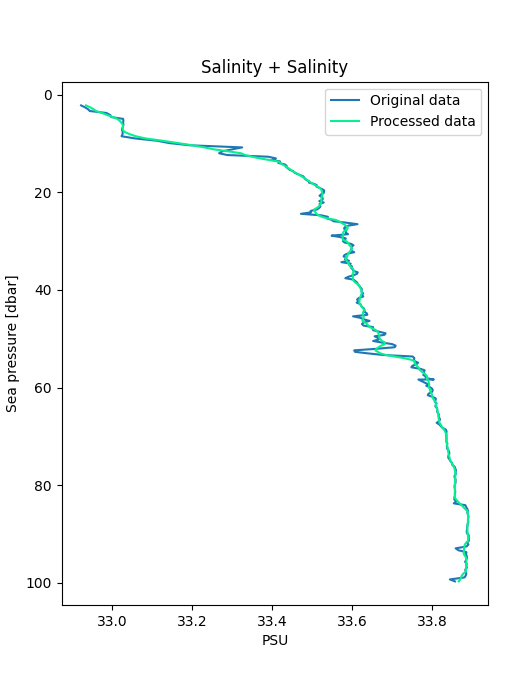
Use RSK.plotprofiles() to compare the binned data to the raw data for a few example profiles.
Processed data are represented with thicker lines.
fig1, axes1 = rsk.plotprofiles(channels=["salinity"],profiles=range(1),direction="down")
rsk.binaverage(binSize = 5, boundary = 0.5, direction = "down")
fig2, axes2 = rsk.plotprofiles(channels=["salinity"],profiles=range(1),direction="down")
fig, axes = rsk.mergeplots(
[fig1,axes1],
[fig2,axes2],
)
for ax in axes:
line = ax.get_lines()[-1]
plt.setp(line, linewidth=0.5, marker = "o", markerfacecolor = "w")
plt.legend(labels=["Original data","Processed data"])
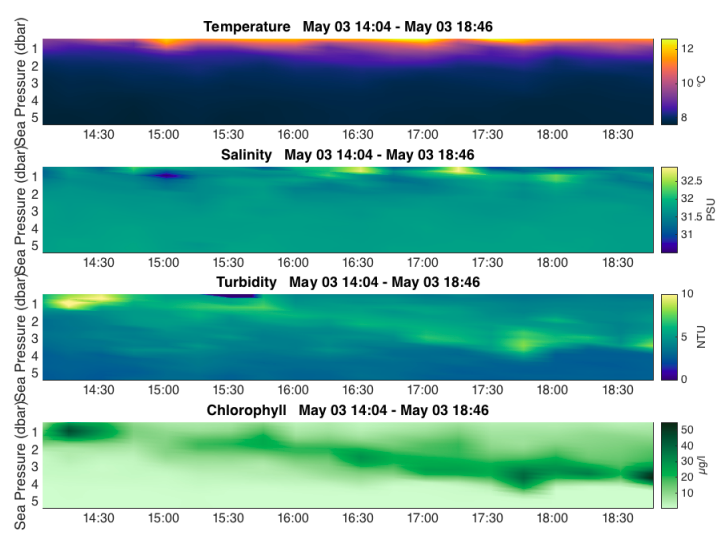
Visualize data with a 2D plot¶
RSK.images() generates a time/depth heat-map of a channel. The x-axis is time; the y-axis is a reference
channel (default is sea pressure). All the profiles must be evaluated on the same reference channel
grid, which is accomplished with RSK.binaverage(). The method supports customizable rendering to
determine the length of the gap shown on the plot. For more details, see RSK.binaverage().
fig,axes = rsk.images(channels= ["temperature","salinity","turbidity","chlorophyll"],direction="up")
Export logger data to CSV files¶
RSK.RSK2CSV() writes logger data and metadata to one or more CSV files. The CSV files contain a header
with important logger metadata, a record of the processing steps made by
pyRSKtools. The data table starts with a row of variable names and units above each column of channel data.
If the profiles number is specified as an argument, then one file will be written for each profile.
Furthermore, an extra column called “cast_direction” will be included. The column will contain ‘d’ or ‘u’
to indicate whether the sample is part of the downcast or upcast, respectively. Users can select
which channels and profiles to write, the output directory, and
also specify additional comments to be placed after the metadata in the file header.
rsk.RSK2CSV(channels = ["depth","temperature","salinity","dissolved_o2_saturation"], profiles= range(0,3), comment= "Hey Jude")
pyRSKtools also has an export method to write Ruskin RSK files (see RSK.RSK2RSK()).
Display a summary of all the processing steps¶
print(rsk.logs)
Other Resources¶
We recommend reading the:
pyRSKtools API reference for detailed pyRSKtools method documentation.
pyRSKtools getting started guide for an introduction on how to make a connection to an RSK file via an instantiated
RSKclass object., make plots, and access the data.
About this document¶
The full source code for this documentation is located at the base of the repository in the docs directory. All documentation is built and generated using Sphinx. For further information, please see Documentation contributions
